|
A. Abstract The Poultry Research Center Virus II (PRC II) is a replication-defective avian sarcoma virus with envelope determinants of the A and B subgroups. In nonproducing cells transformed by PRCII the products of the replicative genes gag, pol, and env are not demonstrable, but a single polyprotein of Mr 105,000 (p105) can be detected. PI 05 contains peptides of the gag proteins p19 and p27 plus transformationspecific sequences. It does not contain peptides of gPr95env of Pr180gag-pol (with the possible exception of one pol peptide) .The transforma tion-specific sequences of p105 are distinct form those of p100 of avian carcinoma virus MH2, of pl10 coded for by avian myelocytoma virus MC29, and of p75 or p40 of avian erythroblastosis virus AEV. They also show no resemblance to p60src of Rous sarcoma virus. P105 is phosphorylated on a tyrosine residue and has an associated phosphokinase activity. P105 appears to be capable of autophosphorylation and of phosphorylating homologous immunoglobulin.
Until recently all avian sarcoma viruses were thought to have fundamentally the same genetic structure. The investigated strains, chiefly Rous sarcoma virus (RSV) and its relatives, contain an about 1.8-kilobase (kb) insertion at the juncture of the env gene and the C region (Duesberg and Vogt 1970; Wang et al. 1975). This transformation-specific insertion, termed src, was found to be of cellular origin and to code for a protein kinase of Mr 60,000 with preference for tyrosine as a phosphoacceptor (Stehelin et al. 1976; Spector et al. 1978 ; Brugge and Erikson 1977; Collett and Erikson 1978; Levinson et al. 1978; Hunter and Sefton 1980). In contrast to RSV, avian acute leukemia viruses code for polyproteins that are derived from a gene in which gag sequences at the 5 I terminal part of the genome are fused to transformation-specific, cell-derived sequences in the middle of the genome. The cell-derived sequences form a substitution for replicative information of the gag, po" and env genes (Bister et al. 1977; Hu et al. 1979; Lai et al. 1979; Sheiness and Bishop 1979; Roussel et al. 1979; Sheiness et al. 1980). As a consequence of this missing information, the acute avian leukemia viruses are replication defective (Ishizaki and Shimizu 1970; Graf 1975 ; Bister and Vogt 1978; Hu and Vogt 1979). A genome organization similar to that of avian acute leukemia viruses has also been found in Abelson and radiation leukemia virus of mice and in feline sarcoma virus (Witte et al. 1978 ; Manteuil- Brutlag et al. 1980; Barbacid et al. 1980; Sherr et al. 1980; Van de Ven et al. 1980). Recently a new class of avian sarcoma viruses has been described which lack src and have agenetic structure related to the avian acute leukemia viruses (Breitman et al. 1981 ; Hanafusa et al. 1980; Kawai et al. 1980; Lee et al. 1980; Neil et al. 1981). The viruses belonging to this group are Fujinami sarcoma virus (FSV), Y73, which is a recent field isolate, and PRCII. Even though these viruses have transformation-specific sequences which are different from the src of RSV, their transformation-specific proteins bear some intriguing functional resemblance to p60src. This functional similarity suggests possible common elements in the mechanism of trans formation by RSV and the new avian sarcoma viruses. The present paper will briefly summarize our studies on PRCII as an eample of the new class of avian sarcoma viruses (Breitman et al. 1981; Neil et al. 1981).
PRCII was isolated from a spontaneous me senterial sarcoma in a
chicken and was shown to cause sarcomas which could be distinguished
histologically from the growths induced by RSV (Carr and Campbell
1958). The virus belongs to envelope subgroups A and B (Payne and
Biggs 1966; Duff and Vogt 1969). PRCII causes only sarcomas in chickens,
and compared to RSV, it is a less virulent agent: Of 50 chickens
inoculated with about 104 focus-forming units of virus, only 13
came down with tumors within an observation time of 4 weeks. In
chick embryo fibroblast cultures PRCII induced foci of predominantly
fusiform transformed cells; but no transformation was seen in cultures
of hematopoietic cells derived from chicken bone marrow or chicken
embryo yolk sac. Infectious center tests showed that while a focus
of PRCII -transformed cells could be induced by a single particle,
a focus which also released infectious progeny was the result of
double infection with transforming virus and nontransforming helper
virus. PRCII-transformed cells which failed to release infectious
virus have been isolated. Such nonproducer lines also failed to
produce noninfectious virions, an indication that the replication
defects in PRCII extend into the gag gene (Bister et al. 1977; Bister
and Vogt 1978; Hu et al. 1978; Hu and Vogt 1979). Superinfection
of these nonproducing cells with avian leukosis helper viruses led
to the rescue of infectious PRCII. In order to search for virus-specific
products in PRCII-transformed cells, virus producing and nonproducing
transformed fibroblasts were labeled with [35S]methionine (50 to
20 µCi/ml) and tested in immunoprecipitation using a variety of
antisera. The precipitates were resolved by polyacrylamide gel electro
phoresis and analyzed by autoradiography (Fig. I ). Rabbit sera
prepared against whole virions of the Prague strain of RSV and sera
against the structural gag protein of avian leukosis viruses precipitated
from PRCII transformed cell lysates a virus-specific protein 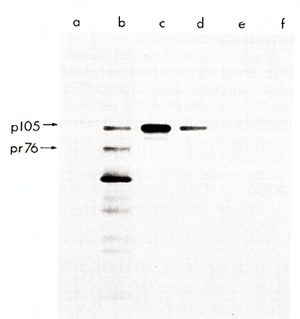
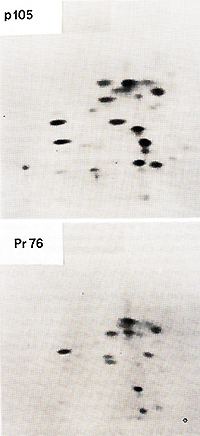
not all of the gagpeptides were present in pI 05 and that pI 05 contained additional peptides not seen in Pr76gag (Fig. 2). Petide maps of the individual gag proteins p19, p27, and p15 were also obtained. The fourth gag protein, p12, does not contain useful [35SJmethionine peptides. These maps of three individual gag proteins showed that all of the p19 and p27 but none of the p 15 peptides were present in pI 05. Identification of the common peptides was confirmed by appropriate mixing experiments. These results lead to the conclusion that p 105 contains the 5 I portion of the gag gene encompassing p19 and p27 but lacks the 3' of gag with p15. No statement can be made concerning the presence or absence of p12 sequences in p105. However, we found a rabbit antiserum prepared against p27 which did not react with p105, although it precipitated p27. Therefore, not all of the determinants of p27 appear to be present in p105. If the order of the gag proteins is N-pI9-p27-pI2-p15-C (Shealy et al. 1980), then p105 may lack p12 and p15 together with a portion from the carboxyterminus of p27. The gag-related part of p105 can then account for roughly half of the p105 molecule. In order to test for the possiblity that the remainder of p105 contains partial, immunologically nonreactive sequencesderived from the pol or env genes, tryptic peptide maps of gPr95env and of Pr180gag-pol of the PRCII helper virus were prepared. There was no overlap between any of the gPr95env peptides and those of pI 05. Of the pol peptides in Pr180, one comigrated with a peptide of p105, and this observation was confirmed in a mixing experiment. However, considering the relative complexity of the PrI80 map, this overlap may be fortuitous and requires further examination. We conclude that p105 does not contain significant portions of the helper virus env or pol genes. Two-dimensional tryptic maps were also obtained of the transformation-specific proteins of (pI10), of carcinoma virus MH2 (p100), of avian erythroblastosis virus AEV strain ES-4 (p75 and p40), and of the p60sre protein from several genetically distinct lines of the Schmidt-Ruppin strain of RSV as well as of the endogenous p60sre of chicken cells. The p60src proteins, the AEV p40, and the non- gag portions of AEV p75, MC29 p110, and MH2 p100 did not contain peptides coinciding with any of p105. We conclude that PRCII contains new transformation-specific sequences not found in RSV or in avian acute leukemia viruses AEV, MC29, or MH2. Cells transformed by PRCII were labeled with (32p]orthophosphate (1 mCi/ml) and followed by analysis by immunoprecipitation with anti-virion sera. Electrophoresis and autoradiography indicated that p 105 was heavily phosphorylated. This degree of phosphorylation was greater than could be accounted for by the presence of the phosphorylated gag protein p19 in pI 05. The most abundant phosphoamino acid in p105 was tyrosine. These observations suggest that pl05 is phosphorylated in its non- gag portion, probably at one or several tyrosine residues. In order to test for phosphokinase activity associated with p105, immunoprecipitates were incubated with (32p]ATP according to published procedures before immunoprecipitation, electrophoresis, and autoradiography. Under these conditions both pl05 and the heavy chain of IgG were phos phorylated. We suggest that p105 is a phosphoprotein with associated kinase activity capable of autophosphorylation and of phosphorylating homologous IgG.
PRCII is a representative of a new class of avian sarcoma viruses that lack src, have a genetic structure similar to avian acute leukemia viruses, and code for a transformation-specific gag-related polyprotein. Other members of this class are FSV and avian sarcoma virus Y73 (Hanafusa et al. 1980 ; Kawai et al. 1980; Lee et al. 1980). FSVand Y73 have been shown to contain a small (26 S) RNA genome. Our preliminary data suggest that the genome of PRCII has also a size of about 26 S. PRCII and FSV habe some common transformation-specific sequences as detected by nucleic acid hybridization (Shibuya et al. 1980), and the proposal has been made to refer to these sequences as Ips (FSV, PRCII Sarcoma) (1. Coffin et al. to be published). Y73 appears to have no appreciable sequence relationship in its transformation-specific in formation to either PRCII, FSV, or RSV (Shibuya et al. 1980; M. Yoshida et al. 1980, personal communications). The transformation-specific sequences of FSV have also been found to be related to those of the SnyderTheilen and Gardner- Arnstein strains of feline sarcoma virus (Shibuya et al. 1980; Barbacid et al. 1981). Since the latter sequences have been detected in normal feline cells where they code for a cellular protein homologue (Barbacid et al. 1981) one can expect Ips to be also cell derived. The transformation-specific polyproteins of FSV, Y73, PRCII, and feline sarcoma virus are all phosphorylated and show phosphokinase activity. The predominant phosphoacceptor for these polyproteins is tyrosine (Hanafusa et al. 1980; Kawai et al. 1980; Neil et al.1981 ; Barbacid et al. 1980). For PRCII and the Snyder- Theilen strain of feline sarcoma virus it has also been shown that phosphotyrosine levels in transformed cells are substantially increased over normal cells (Barbacid et al. 1980; K. Beemon 1980, personal communication). Phosphorylation and kinase activity are also characteristic of p60src, and cells transformed by RSV contain elevated levels of phosphotyrosine ( Collett and Erikson 1978; Levinson et al. 1978; Hunter and Sefton 1980). These parallels suggest that even though sarcoma viruses may carry different transforming genes, the mechanisms of viral sarcomagenesis could share common elements in protein function and cellular targets.
Supported by U.S. Public Health Service Research Grants No. CA 1 3213 and 19725 awarded by the National Cancer Institute.
- Barbacid M, Beemon K, Devare SG (1980) Proc Natl Acad Sci USA
77:5158-5162 |