|
1 Division of Hematology/Oncology, Washington University,
St. Louis, MO
2 Laboratory of Tumor Cell Biology, National Cancer Institute, Bethesda,
MD
3 Biotech Research Laboratories, Inc., Rockville, MD
4 The Clinical Oncology Program, National Cancer Institute, Bethesda
There is considerable evidence that the human T -lymphotropic virus
type III/lymphadenopathy-associated virus (HTL V -III/ LA V) is
the etiological agent in the acquired immunodeficiency syndrome
(AIDS) and AIDS-related syndromes [1]. The most convincing line
of evidence is the recapitulation with in vitro infection of the
major manifestation of the disease, depletion ofT4 cells [2]. Definition
of the viral determinants of this lymphocytopathic activity is critical
to understanding the pathogenesis of AIDS. With this goal in mind
we have established an in vitro model which will facilitate this
analysis. In this system, plasmid clones with the full HTLV-III/LA
V proviral sequence are transfected into normal human umbilical
cord blood mononuclear cell cultures. This results in production
of virus particles with a morphology typical of HTL V -III/LA V
and in death of the cell culture (Fig. 1). To provide the basic
information for utilization of this assay system, we have determined
the complete nucleotide sequence of the biologically active proviral
clone HXB2 [3]. Eighty nucleotide substitutions are noted, compared
to the previously reported HTLV-III/LAV sequence for clone BH10
[4]. Insertions of two and three nucleotides in HXB2 compared to
BH10 were recognized in noncoding regions, as well as a deletion
of one copy of a 36-nucleotide, tandemly repeated sequence in the
overlap of gag and pol. Most notable is the lack of alterations
in the size and location of each of the seven previously identified
viral genes [3]. Polymorphism is also noted in the predicted amino
acid sequences of the viral protein products of HXB2 compared to
the other sequenced HTL V -III/LA V viruses. Gag, pol, and sor are
relatively well conserved, with 0.6%-3.6%, 1.0%-4.0%, and 0.5%-10.9%
amino acid substitutions respectively. Tat, trs, env, and 3'orjare
more polymorphic, with 0.0%-11.6%, 5.2%16.3%, 1.7%-17.5%, and 2.9%-16.0%
amino acid substitutions respectively. A number of amino acid insertions
and deletions are also noted. The relationships of these sequence
variations to alterations in neutralizing epitopes, receptor binding
domains, and other biological characteristics of the virus remain
to be determined. The use of molecularly cloned viruses generated
from this in vitro system will provide reagents for approaching
these problems. The biological activity of several other HTL V -III/LA
V clones was also tested in umbilical cord blood mononuclear cells
and the T4 + cell line, A TH8 [3]. To test the functional capabilities
of clone BH10, the missing portions of the provirus were complemented
with long-terminal repeat sequences from HXB2. The resultant clone
gave rise to a lymphocytopathic virus. Clone HXB3 has been partially
sequenced and appears to be closely related to HXB2, differing at
only 63 of 3890 positions in the 3' portion of the genome. A notable
difference between HXB2 and HXB3, however, is the presence of a
termination codon in 3' orf of HXB2 and the lack of this sequence
in HXB3. Viruses generated from HXB2 and HXB3 have similar replicative
and cytopathic abilities. This finding, together with additional
data on clones with deletions in 3'orf(see Fisher et al., this volume),
suggests that 3' orfplays no essential role in the ability of HTL
V -III/LA V either to replicate or to kill T4-lymphocytes.
Thus, these data provide the basic information essential for utilization
of this system to construct clones of HTL V -III/LA V with alterations
in the viral genome and for their assay in human lymphoid cells.
Application of this system has yielded a variant with markedly attenuated
cytopathic activities but normal replication (see Fisher et al.,
this volume). Further applications should provide information essential
to understanding the pathogenesis of cell killing in AIDS and lead
to approaches to the treatment and prevention of this disease.
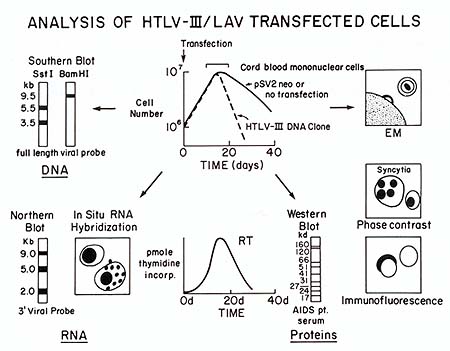
Fig. 1. An in vitro model system for AIDS. The schematic
drawing depicts the major characteristics of the transfection system
for analysis of the biological activity of HTL V -III/LA V DNA clones.
The top-center drawing shows the growth curve of umbilical cord
blood mononuclear cells after transfection with a plasmid lacking
HTL V III/LA V DNA sequences (pSV2neo) or a plasmid with the full
HTL V -III/LA V provirus. Parameters measured 7-25 days after transfection
include analysis of viral DNA by Southern blot, of viral RNA by
Northern blot or in situ hybridization, of viral proteins by reverse
transcriptase assays, Western blot, immunofluorescence for gag or
env products, or phase-contrast microscopy for syncytia formation,
and of the expression of particles with a morphology characteristic
of HTL V III/LA V
References
1. Gallo RC, Wong-Staal F (1985) A human Tlymphotropic retrovirus
(HTL V -III) as the cause of the acquired immunodeficiency syndrome.
Ann Int Med 103:679-689
2. Popovic M, Sarngadharan MG, Read E, Gallo RC (1984) Detection,
isolation, and continuous production of cytopathic retroviruses
(HTLV-III) from patients with AIDS and preAIDS. Science 224:497-500
3. Ratner L, Fisher A, Jagodzinski LJ, Mitsuya H, Liou R-S, Gallo
RC, Wong-Staal F (1987) Complete nucleotide sequences of functional
clones of the virus associated with the acquired immunodeficiency
syndrome, HTL V -III/
LA V. AIDS Res & Hum Retrov 3
4. Ratner L, Haseltine W, Patarca R, livak KJ , Starcich B, Josephs
SF, Doran ER, Rafalski JA, Whitehorn EA, Baumeister K, Ivanoff L,
Petteway SR, Pearson ML, Lautenberger JA, Papas TS, Ghrayeb J, Chang
NT, Gallo RC, Wong-Staal F (1985) Complete nucleotide sequence of
the AIDS virus, HTLV-III. Nature 313:277-284
|
