Transduction of myc and a T -Cell Antigen Receptor ß-Chain Gene |
|
1 Beatson Institute for Cancer Research, Bearsden,
Glasgow
Feline leukaemia virus has been a particularly useful tool in cancer research since many of the naturally occurring tumours associated with this virus group have yielded recombinant retroviruses containing hostderived oncogenic information. The prevalence of transduction as an oncogenic mechanism was seen first in multicentric fibrosarcoma, a relatively rare tumour in FeL V -infected cats (Hardy et al. 1982; Besmer 1984). In a significant percentage of cases of this disease, oncogene-containing feline sarcoma viruses have been identified. More recently, we and others have found that in the more common FeL V -associated neoplasm, thymic lymphosarcoma, viral capture of the c-myc gene can occur (Neil et al. 1984; Levy et al. 1984; Mullins et al. 1984). Since the oncogenes carried by feline sarcoma viruses do not include myc, these reports provided the first evidence that myc may be a target for oncogenic activation by FeL V. Further study of feline tumours revealed that c-myc could be affected either by viral transduction or by proviral insertion into the cellular gene locus (Neil et al. 1984, 1987; Forrest et al. 1987) although the majority of field-case tumours showed neither of these features. To gain more information on the cellular origin of the feline lymphoid tumours and to search for some distinguishing feature of those with activated myc genes, we undertook an analysis of the state of rearrangement and expression of the genes encoding IX and ß chains of the T -cell antigen receptor. In the course of this analysis we discovered a novel FeL V provirus in which a full-length ß-chain gene had been incorporated into the viral genome (Fulton et al. 1987). The present report describes this novel provirus and considers its possible significance in leukaemogenesis.
Analysis of feline T -cell receptor genes was performed with cDNA probes derived from the human IX (pJIX6) and ß (pB400) genes (Collins et al. 1985 b); these were kindly provided by Michael Owen (ICRF Tumour Immunology Labs, London). We found these probes to be strongly cross-reactive with the feline genes and their transcripts, although hybridisation was stronger with the ß-chain than with the IX-chain probe. For this reason, only the ß-chain probe has been used to assess gene rearrangement. The overall gene arrangement and transcript sizes appear similar for feline and human genes. Thus, as in the human and the murine Cß loci (Gascoigne et al. 1984; Malissen et al. 1984; Sims et al. 1984) the feline Cß coding sequence appears to be tandemly duplicated in germ-Iine DNA. Also, some of the feline tumours examined displayed rearrangement of both Cß alleles with two distinct transcript sizes of 1.2 and 1.4 kb, which by analogy with human genes (Collins et al. 1985a) Table I. Rearrangements of T -cell
antigen receptor ß-chain gene in feline leukaemias 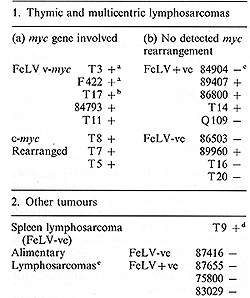
correspond to abortive (D-J-C) and successful (V-D-J-C) joining events, respectively. The results of a survey of ß-chain rearrangement are given in Table 1. The conclusions both from these data and from those of Northern blot analyses for the expression of alfa- and ß-chain transcripts are that feline thymic tumours are heterogeneous with respect to maturity as assessed by T -cell receptor gene rearrangement and expression. However, the tumours involving cmyc activation, either by transduction or by proviral insertion, represent a homogeneous subset with mature characteristics (expressing both alfa- and ß-chain transcripts).
Tumour T17 showed an anomalous pattern of ß-chain mRNA both in
size (>6 kb) and in abundance. Furthermore, DNA blots showed gross
amplification of sequences hybridising to the human Cß probe. Further
Southern blot hybridisation analysis of tumour T17 showed that the
amplified sequences could be resolved into a single, intensely hybridising
fragment if digestion was performed with any of the enzymes which
characterise the FeLV LTR (Kpnl, Smal, Pstl and Hincll) (Fig. 1).
These data provide indirect but persuasive evidence that the amplification
of ß-chain sequences in tumour T17 was due to their presence within
multiple FeLV proviruses. Cloning was undertaken to isolate the
novel proviral structures from tumour T17. From a library of size
selected (15-23 kb), EcoRI-digested tumour DNA in lambda EMBL 4
we selected recombinants with various probes, including the human
Cß cDNA clone, FeL V v-myc, FeL V env and FeL V L TR. The clones
we have isolated correspond to the proviruses containing Cß sequences,
proviruses containing v-myc, FeL V helper-type proviruses and the
normal cellular loci of c-myc and Cß. Our initial efforts have focussed
upon characterising the FeL V proviruses containing the Cß-hybridising
sequences. A 1.9-kb fragment containing the entire hybridising sequence
was sequenced and found to contain a 1.2-kb host-derived sequence
insert including the intact coding sequence of a ßchain T -cell
receptor gene (Fulton et al. 1987; see Fig. 1 ). The ß-chain gene
appears to have undergone productive rearrangement since sequences
clearly identifiable as those of V ß, Dß, J ß and Cß origin are
seen. lntron sequences are missing, however, as might be expected
if the sequence has been transmitted as part of a retroviral replication
unit. 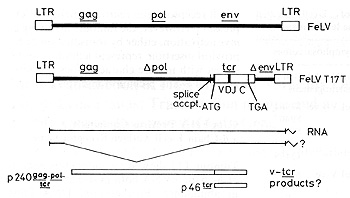
The host-derived sequence replaces the 3' end of pol and much of the FeL V env gene. The recombination junction is 7 base pairs upstream of the A TG, marking the beginning of the ß-chain open reading frame which begins with the characteristic signal peptide for membrane insertion. The 3' end of the host insert appears to be coincident with the polyadenylation signal (AATAAA). In this context the host-derived sequence (which we have designated v-tcr) could be expressed as a protein in two different ways. Firstly, since the pol reading frame is coincident with that of v-tcr, the gene may be expressed as a large fusion protein including gag, pol, and tcr sequences. Alternatively, the proximity of the splice acceptor site for the env mRNA means that a spliced subgenomic RNA could direct the synthesis of a v-tcr product which is neither truncated nor fused to viral sequences (Fig. 1). We are at present investigating these possibilities.
Rather less information is available at present regarding the myc-containing provirus from tumour T17 .Although EMBL 4 phage clones were readily obtained, full-length sub-clones have proved impossible to obtain thus far. Possible "poison" sequences have not yet been located in the provirus or its flanking sequences, but cloning in segments into plasmid vectors has allowed us to isolate possibly all the proviral structure. Since T17 represents the first recorded example of a "double transduction" event in which both host genes are present on separate proviruses, we wish to discover whether these recombinants arose independently or whether one recombinant provirus may have arisen from the other. Initial mapping suggests that the myc gene replaces env, as in the v-tcr-containing virus, however we do not yet know the precise 5' and 3' junctions.
Although FeLV myc recombinant viruses appear to be potent initiators of tumour development, several features led us to consider that the FeL V v-myc genes may be insufficient for full neoplastic development. The first factor we considered is the latent period for FeL V myc virus-induced tumour development, which is shorter than that for helper FeL V but longer than that for many other v-onc-containing retroviruses. Also, analysis of integrated proviruses by Southern blot hybridisation with FeL V or v-myc probes indicated that even the short latency FeL V myc tumours represent monoclonal or oligoclonal outgrowths of virus-infected cells. Furthermore, these tumour cells could be established readily in culture in the absence of exogenous sources of interleukin 2 (IL-2), although such transformed cell lines could not be obtained in vitro even after a series of attempts to infect isolated T cells or bone marrow cell cultures (Onions et al. 1987). While these phenomena may also be explained in other ways, we consider that the sum of the evidence points to secondary oncogenic events in vivo which we cannot so far reproduce in vitro. The finding here of a novel provirus containing a T -cell antigen receptor gene in the same tumour as a v-myc gene suggests a possible secondary oncogenic factor for this one case. In the majority of cases which do not show such proviruses we must seek other explanations. However, the observation that all of the tumours involving direct myc activation are of mature T -cell phenotype may provide a useful clue. The oncogenic properties of v-tcr have not yet been tested by in vivo experiments. The primary tumour from T17 is no longer available and was in any case a very poor virus producer. We have therefore had to resort to transfection experiments to reconstruct virus complexes for inoculation into cats; these experiments are in progress. Predictably perhaps, initial experiments have shown no transforming potential of v-tcr for fibroblastic cells. Transfections into mature human T cells have been undertaken to discover whether the v-tcr gene product(s) can interact with human alfa chain and other T -cell receptor components and lead to membrane transport of the complex. At the same time, we will monitor any disturbance in growth or responsiveness to external stimuli (e.g. lectins, phorbol esters) which may give clues to the mode of action of v-tcr. Our initial hypothesis was that v-tcr might cause constitutive activation of the antigen receptor in the absence of external antigen (Fulton et al. 1987). The rationale for this model was that the transmembrane region of v-tcr has a nonconservative change (met > Iys) relative to the human and mouse Cß sequences. Thus, in a manner akin to the proposed mechanism of activation of the neu gene (Bargmann et al. 1986), altered conformation might mimic the presence of extracellular ligand. This model now appears less likely for v-tcr in view of our finding that the cellular Cß locus cloned from tumour T17 has the same transmembrane sequence as the viral gene (1 .N .and R.F ., unpublished results). The difference which we recorded (Fulton et al. 1987) may therefore be a species-specific change. These results leave both the oncogenic significance and the possible modes of action of v-tcr as open questions. We will have to await the outcome of in vivo experiments for the answer to the first question. We may then have to address the possibility that the immunological specificity carried by v-tcr is the key to its oncogenic function. Recognition of a host or viral antigen seems possible, although the associated alfa chain might then be expected to playa contributory role. If self-reactivity is involved, we may speculate further that the v-tcr specificity would normally have been suppressed during thymic education of lymphocytes where specificity for host-MHC plus foreign antigen is learned (Bevan 1981). This self-reactivity may have been augmented in T17 by the retroviral capture of the important part of the immune effector involved in recognition. We might also propose that the role ofv-m.yc in this case is to rescue (immortalise) a cell clone with autologous reactivity and "selfdriven" proliferative capacity. Whatever the explanation, it is our hope that v-tcr may contribute in a wider sense to our understanding of normal and neoplastic T -cell growth. Acknowledgements.
1. Bargmann CI, 1Hung M-C, Weinberg RA (1986) Multiple independent
activations of the ncu oncogene by a point mutation altering the
transmembrane domain of p185. Cell 45.649-657 |